A linkage group is a group of genes whose loci are on the same chromosome and hence don’t independently assort
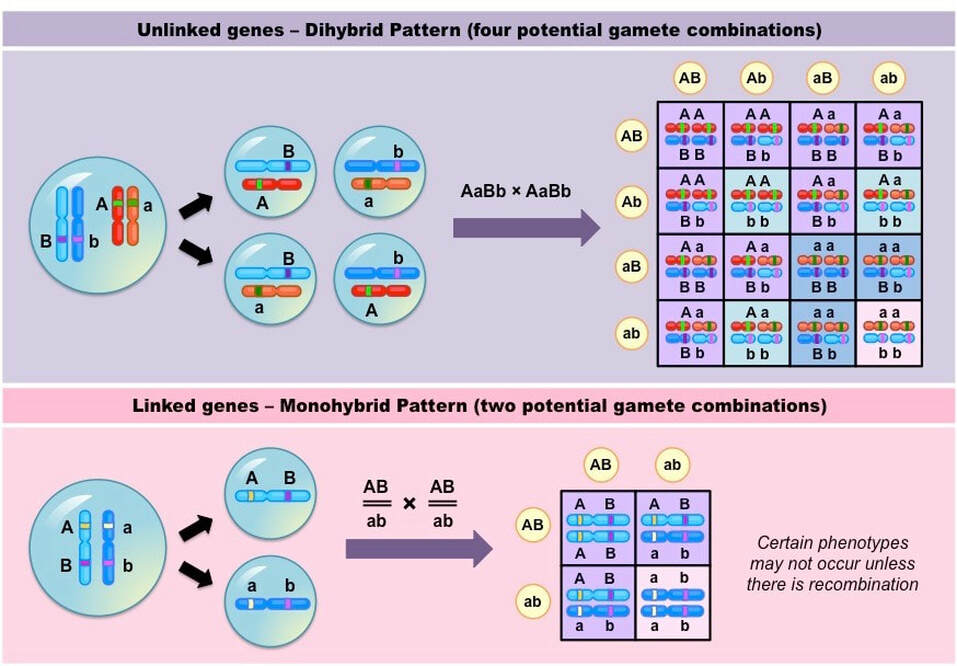
- Linked genes will tend to be inherited together and hence don’t follow normal Mendelian inheritance for a dihybrid cross
- Instead the phenotypic ratio will be more closely aligned to a monohybrid cross as the two genes are inherited as a single unit
- Linked genes may become separated via recombination (due to crossing over during synapsis in meiosis I)
Unlinked versus Linked Inheritance Patterns
Application:
10.2.A2: Morgans’s discovery of non-Mendellian ratios in Drosophila
Objectives:
10.2.A2: Morgans’s discovery of non-Mendellian ratios in Drosophila
Objectives:
- Describe how Morgan discovered relationship between eye color and sex in Drosophila.
Thomas Hunt Morgan provided a key contribution to our current understanding of gene linkage by discovering non-Mendelian ratios in Drosophila melanogaster (fruit flies)
Sex Linkage
When cross-breeding red-eyed wild types with white-eyed mutants, he discovered a clear sex bias in phenotypic distribution
- His breeding experiments involving fruit flies clearly demonstrated that linked genes were not independently assorted
Sex Linkage
When cross-breeding red-eyed wild types with white-eyed mutants, he discovered a clear sex bias in phenotypic distribution
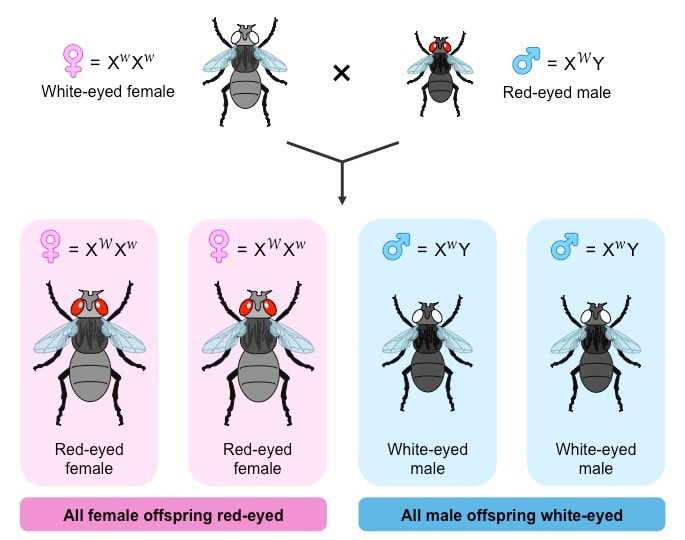
- All female offspring of a red-eyed male were red-eyed, whereas all male offspring of a white-eyed female were also white-eyed
- Morgan described this distribution as 'sex-limited’ inheritance and inferred it was caused by the gene for eye colour being located on a sex chromosome (i.e. X-linked)
Morgan's Discovery of Sex Linkage in Drosophila
Gene Linkage
Morgan went on to identify a number of different traits in fruit flies that did not conform to Mendelian ratios
Based on this data, Morgan made two key proposals:
Morgan also observed that the amount of crossing over between linked genes differed depending on the combination of traits
Morgan went on to identify a number of different traits in fruit flies that did not conform to Mendelian ratios
- Certain phenotypic combinations occurred in much lower frequencies than was to be expected
Based on this data, Morgan made two key proposals:
- The alleles for these traits were located on a shared chromosome (gene linkage) and hence did not independently assort
- Linked alleles could be uncoupled via recombination (crossing over) to create alternative phenotypic combinations, but these new phenotypes would occur at a much lower frequency
Morgan also observed that the amount of crossing over between linked genes differed depending on the combination of traits
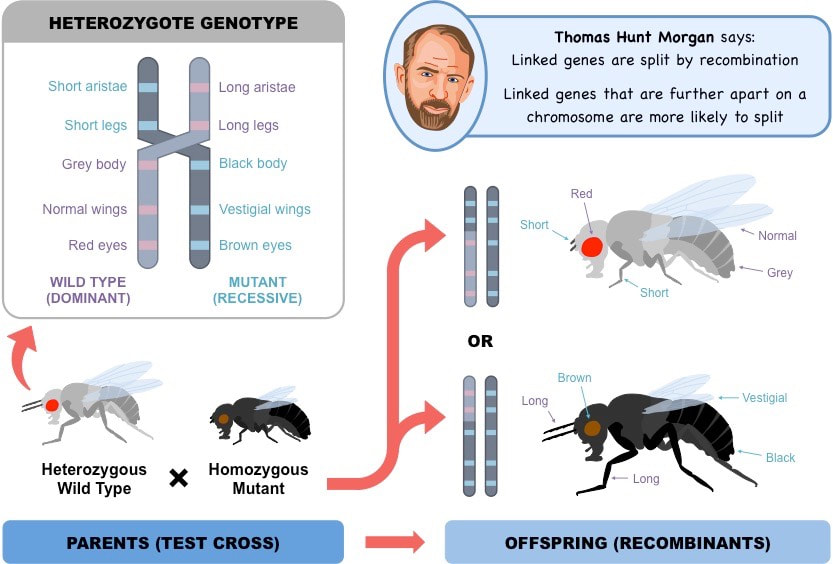
- This led to the idea that crossover frequency may be a product of the distance between two genes on a chromosome – genes with a higher crossover frequency are further apart, whereas genes with a lower crossover frequency are closer together
- Morgan used this concept to develop the first gene linkage maps that showed the relative positions of genes on a chromosome
Gene Linkage and Recombination Frequencies
Skills:
10.2.S2: Identification of recombinants in crosses involving two linked genes
Objectives:
10.2.S2: Identification of recombinants in crosses involving two linked genes
Objectives:
- Use correct notation to show alleles of linked genes.
- Construct a Punnett square to show the possible genotype and phenotype outcomes in a dihybrid cross involving linked genes.
- Explain how crossing over between linked genes can lead to genetic recombinants.
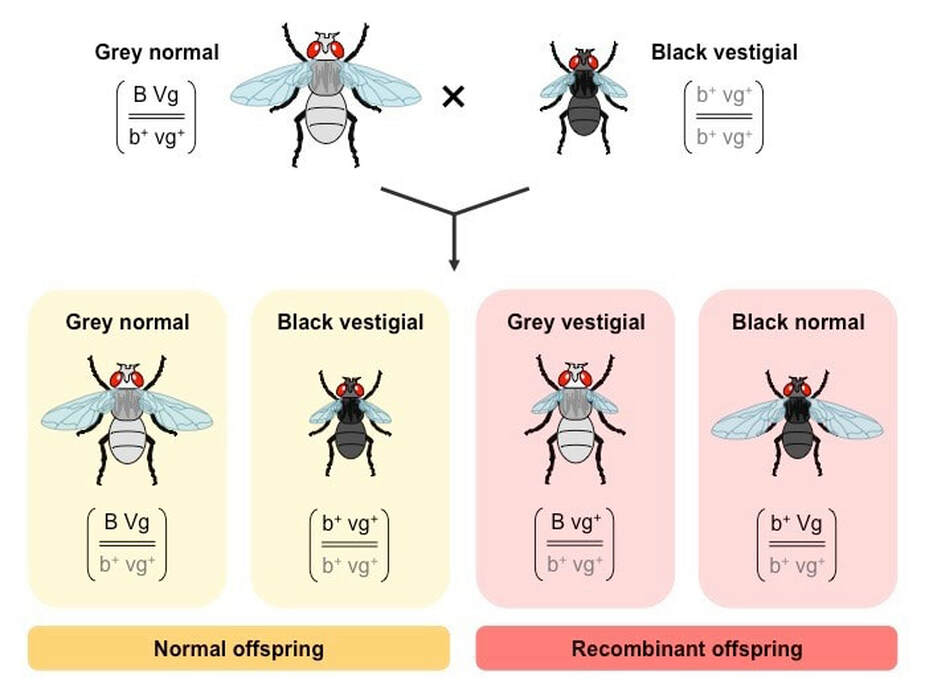
Recombinants of linked genes are those combinations of genes not found in the parents
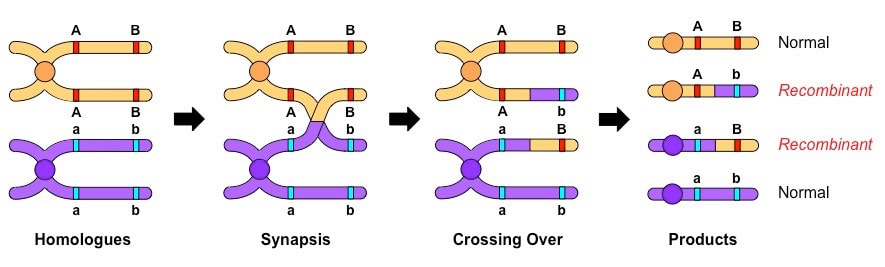
- Recombinants occur as a result of crossing over of genetic material during prophase I of meiosis
- If linked genes become separated by a chiasma, there will be an exchange of alleles between the non-sister chromatids
- This creates new allele combinations that are different to those of the parent
Identifying Recombinants from Genotype
The frequency of recombinant phenotypes within a population will typically be lower than that of non-recombinant phenotypes
The relative frequency of recombinant phenotypes will be dependent on the distance between linked genes
Recombinant phenotypes can be identified by performing a test cross (crossing with a homozygous recessive for both traits)
- This is because crossing over is a random process and chiasmata do not form at the same locations with every meiotic division
The relative frequency of recombinant phenotypes will be dependent on the distance between linked genes
- Recombination frequency between two linked genes will be greater when the genes are further apart on the chromosome
- This is because there are more possible locations where a chiasma could form between the genes
Recombinant phenotypes can be identified by performing a test cross (crossing with a homozygous recessive for both traits)
Identifying Recombinants from Phenotype