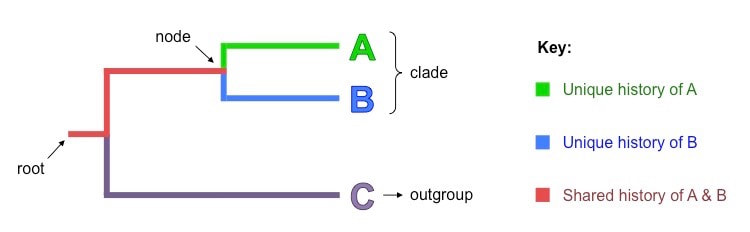
Constructed cladograms all typically share certain key features:
- Root – The initial ancestor common to all organisms within the cladogram (incoming line shows it originates from a larger clade)
- Nodes – Each node corresponds to a hypothetical common ancestor that speciated to give rise to two (or more) daughter taxa
- Outgroup – The most distantly related species in the cladogram which functions as a point of comparison and reference group
- Clades – A common ancestor and all of its descendants (i.e. a node and all of its connected branches)
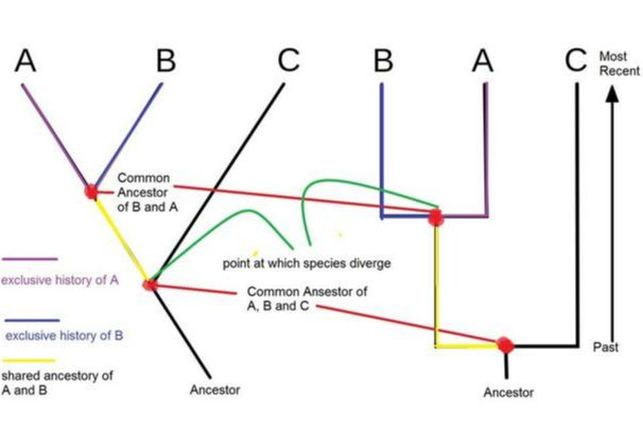
Analyzing Cladogram Practice
|
The red dots are called nodes, and represent the time when two species are estimated to have split.
One thing to note, just because a species like C split earlier than from B, it does not mean that B has evolved more. All the species at the top are present D species. The ones that have died out or changed would be at the nodes. |
Constructing Cladograms
Cladograms can be constructed based on either a comparison of morphological (structural) features or molecular evidence
1. Using Structural Evidence
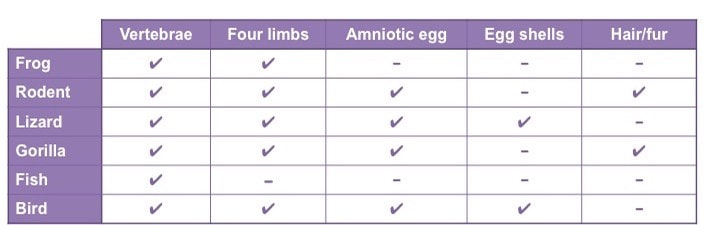
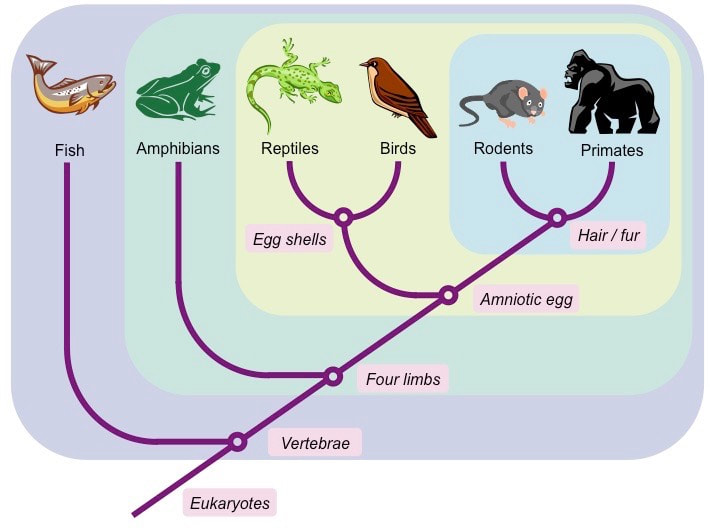
Step 1: Organise selected organisms according to defined characteristics
Cladograms can be constructed based on either a comparison of morphological (structural) features or molecular evidence
- Historically, structural features were used to construct cladograms, but molecular evidence is now more commonly used
1. Using Structural Evidence
Step 1: Organise selected organisms according to defined characteristics
- Use characteristics that are developmentally fixed (i.e. innate) and not influenced by environmental pressures
Constructing Cladograms
Cladograms can be constructed based on either a comparison of morphological (structural) features or molecular evidence
Historically, structural features were used to construct cladograms, but molecular evidence is now more commonly used
1. Using Structural Evidence
Step 1: Organise selected organisms according to defined characteristics
Use characteristics that are developmentally fixed (i.e. innate) and not influenced by environmental pressures
Cladograms can be constructed based on either a comparison of morphological (structural) features or molecular evidence
Historically, structural features were used to construct cladograms, but molecular evidence is now more commonly used
1. Using Structural Evidence
Step 1: Organise selected organisms according to defined characteristics
Use characteristics that are developmentally fixed (i.e. innate) and not influenced by environmental pressures
2. Using Molecular Evidence
Step 1: Select a gene or protein common to a range of selected organisms
Examples of molecules which are ubiquitously found in many animals include haemoglobin and cytochrome c
Step 2: Copy the molecular sequence (DNA or amino acid) for each of the selected organisms
Use online databases such as Genbank or Ensembl to identify relevant DNA or amino acid sequences
Sequences can be collated in a Word document and then saved as a document in plain text format (.txt)
Before each sequence, designate a species name preceded by a forward arrow (e.g. '>Human’ or ‘>Chimpanzee’)
Step 3: Run a multiple alignment to compare molecular sequences (DNA or amino acid)
Multiple alignment software compares DNA or protein sequences for similarities and differences
Closely related species are expected to have a higher degree of similarity in their molecular sequence
Clustal Omega is a free online tool that will align multiple DNA or amino acid sequences for comparison
Step 4: Generate a phylogeny tree (cladogram) from multiple alignment data
Clustal Omega can generate branched phylograms after a sequence alignment is completed (select ‘Phylogenetic Tree’)
Below is a plain text file that can be uploaded to compare amino acid sequences from different species:
HBA – Haemoglobin alpha chain (amino acid sequence) from various species
Step 1: Select a gene or protein common to a range of selected organisms
Examples of molecules which are ubiquitously found in many animals include haemoglobin and cytochrome c
Step 2: Copy the molecular sequence (DNA or amino acid) for each of the selected organisms
Use online databases such as Genbank or Ensembl to identify relevant DNA or amino acid sequences
Sequences can be collated in a Word document and then saved as a document in plain text format (.txt)
Before each sequence, designate a species name preceded by a forward arrow (e.g. '>Human’ or ‘>Chimpanzee’)
Step 3: Run a multiple alignment to compare molecular sequences (DNA or amino acid)
Multiple alignment software compares DNA or protein sequences for similarities and differences
Closely related species are expected to have a higher degree of similarity in their molecular sequence
Clustal Omega is a free online tool that will align multiple DNA or amino acid sequences for comparison
Step 4: Generate a phylogeny tree (cladogram) from multiple alignment data
Clustal Omega can generate branched phylograms after a sequence alignment is completed (select ‘Phylogenetic Tree’)
Below is a plain text file that can be uploaded to compare amino acid sequences from different species:
HBA – Haemoglobin alpha chain (amino acid sequence) from various species
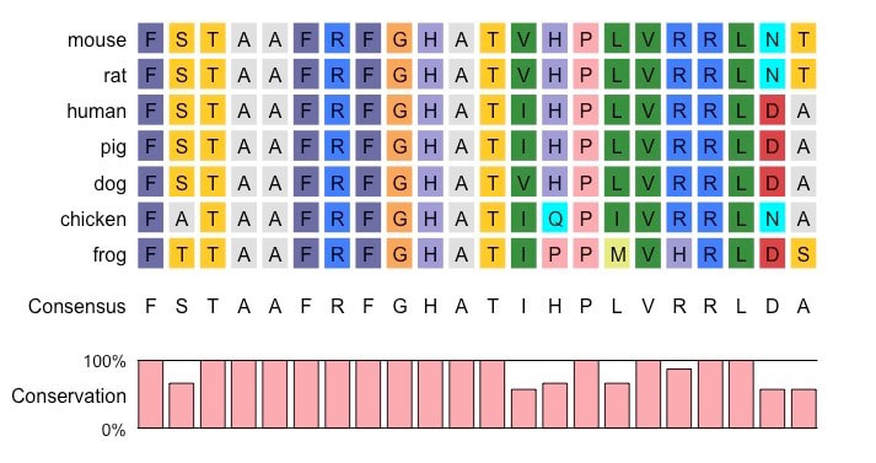
Multiple Alignment of a Protein Sequence from Various Species